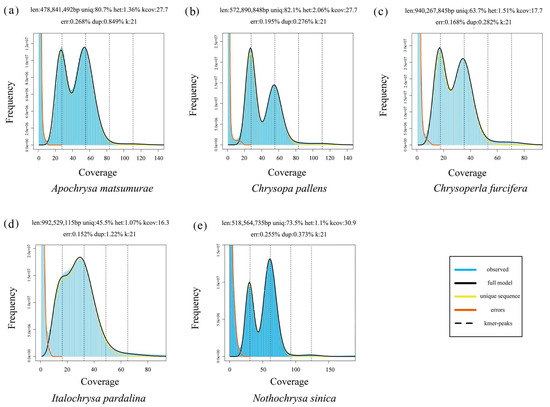
Analysis of population structure and genetic diversity in low-variance Saimaa ringed seals using low-coverage whole-genome sequence data
By A Mystery Man Writer
Description
STAR Protocols is an open access, peer-reviewed journal from Cell Press. We offer structured, transparent, accessible, and repeatable step-by-step experimental and computational protocols from all areas of life, health, earth and physical sciences.

Fragmented habitat compensates for the adverse effects of genetic

A) Geographic distributions of the three northern European ringed

Map of study areas of radio-tagged Saimaa ringed seals: (1) Sulevi, (2)

Whole-genome resequencing reveals genetic diversity and selection
Population-level genome-wide STR discovery and validation for

Ari LÖYTYNOJA, PhD, University of Helsinki, Helsinki, HY, Institute of Biotechnology

Integrative Evolutionary Biology

Analysis of population structure and genetic diversity in low

Ari LÖYTYNOJA, PhD, University of Helsinki, Helsinki, HY, Institute of Biotechnology
from
per adult (price varies by group size)